'szip' may be unavailable as it depends on the HDF5 installation including it. Extra saving arguments Ĭompression: One of None, 'gzip', 'szip', 'lzf' (default is 'gzip'). See the chunking section for more information. 
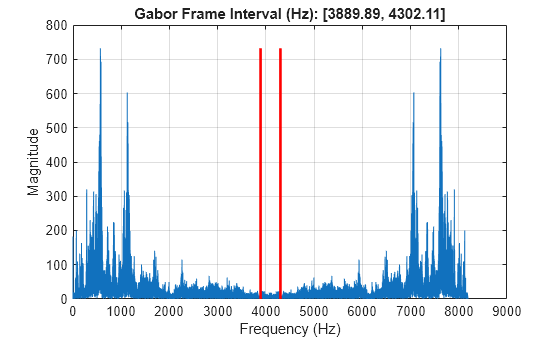
Passing chunks=True results in (7, 7, 256) chunks.Ĭhoosing the correct chunk-size can significantly affect the speed of reading, writing and performance of many HyperSpy algorithms. For example, for the signal in the example above What, for large signal spaces usually leads to smaller chunks as guess_chunk does not impose theĬonstrain of storing at least one signal per chunks. Will be applied to all arrays contained in the signal.īy passing True to chunks the chunk shape is guessed using h5py’s guess_chunk function Therefore, the value of chunks provided on saving Note that currently it is not possible to pass different customised chunk shapes to all signals andĪrrays contained in a signal and its metadata. Part of the specification is documented in Metadata structure. The HDF5 file format is supported by many Information will be lost in the writing process and that supports saving data This is the default format and it is the only one that guarantees that no The “lazy” column specifies if lazy evaluation is supported. Here is a summary of the different formats that are currently supported by The following will write the data to a file called spectrum.hspy in theĭefault HSpy - HyperSpy’s HDF5 Specification format: For example, if the s variableĬontains the BaseSignal that you want to write to a file, ( HSpy - HyperSpy’s HDF5 Specification) is used. If the filename does not contain the extension, the default format Theįirst argument is the filename and the format is defined by the filenameĮxtension. To save data to a file use the save() method. load ( '*.rpl', stack = True ) > s Saving data to files > ls CL1.raw CL1.rpl CL2.raw CL2.rpl C元.raw C元.rpl CL4.raw CL4.rpl L元.raw L元.rpl shift_map-SI3.npy hdf5/ > s = hs. Tips for writing methods that work on lazy signals.HDF5 reader plugin for Digital Micrograph.Reading data generated by HyperSpy using other software packages.Extra loading arguments for SPD and SPC files.Energy-Dispersive X-ray Spectrometry (EDS).Interactive Operations and Region of Interest (ROI). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
